MSc Thesis | Delft University of Technology | Grade: 9.5
Author: Gijs Bernd Schlief
Welcome to the repository for my Msc thesis on Using Reinforcement Learning to Solve the Inverse Bone Remodelling Problem. This repository contains all the code, scripts, and resources developed and used during the research. Special thanks are due to Bansod, whose code was a major source of inspiration for the forward model implementation. Please do check out his implementation on: https://github.com/YDBansod/Bone_Remodelling. A GIF demonstrating the forward model's capabilities is shown below!

Figure 1: The forward model adapting over time using a standard triangular loading condition on the top
The agent's reward function is designed to bridge the gap between observed morphology and mechanical history. By penalising the Mean Squared Error (MSE) between the target bone density and the simulated density, the PPO (Proximal Policy Optimisation) agent learns to navigate a high-dimensional search space that is traditionally prone to local minima. As shown in Figure 2, the reward curve demonstrates stable convergence over 20 million timesteps. The agent successfully transitions from exploratory "random" loading to an exploitation phase, where it identifies the specific force vectors that replicate the target structure with high fidelity.
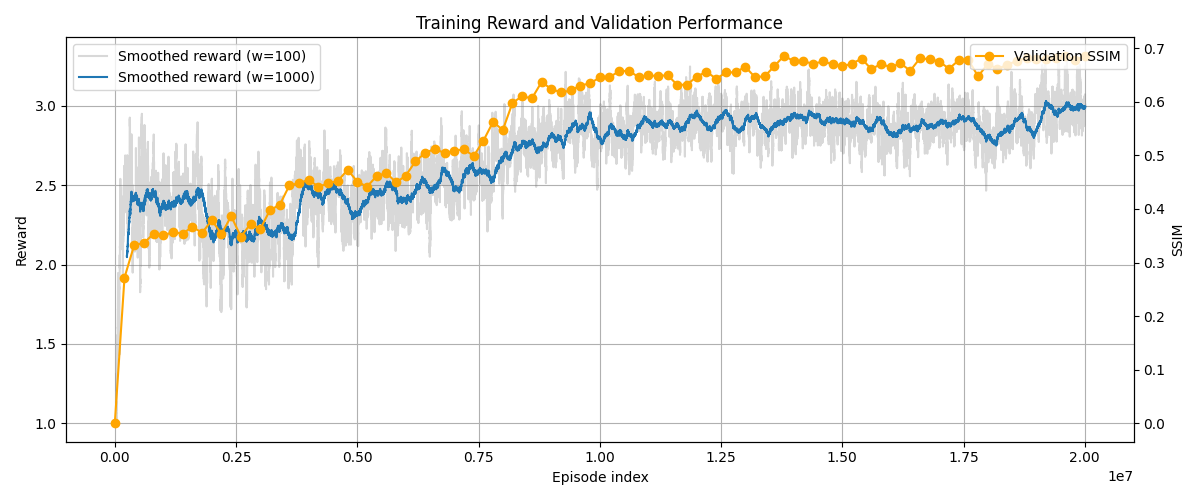
Figure 2: The forward model adapting over time using a standard triangular loading condition on the top
The core challenge of this thesis was to outperform traditional supervised learning and gradient-based methods.
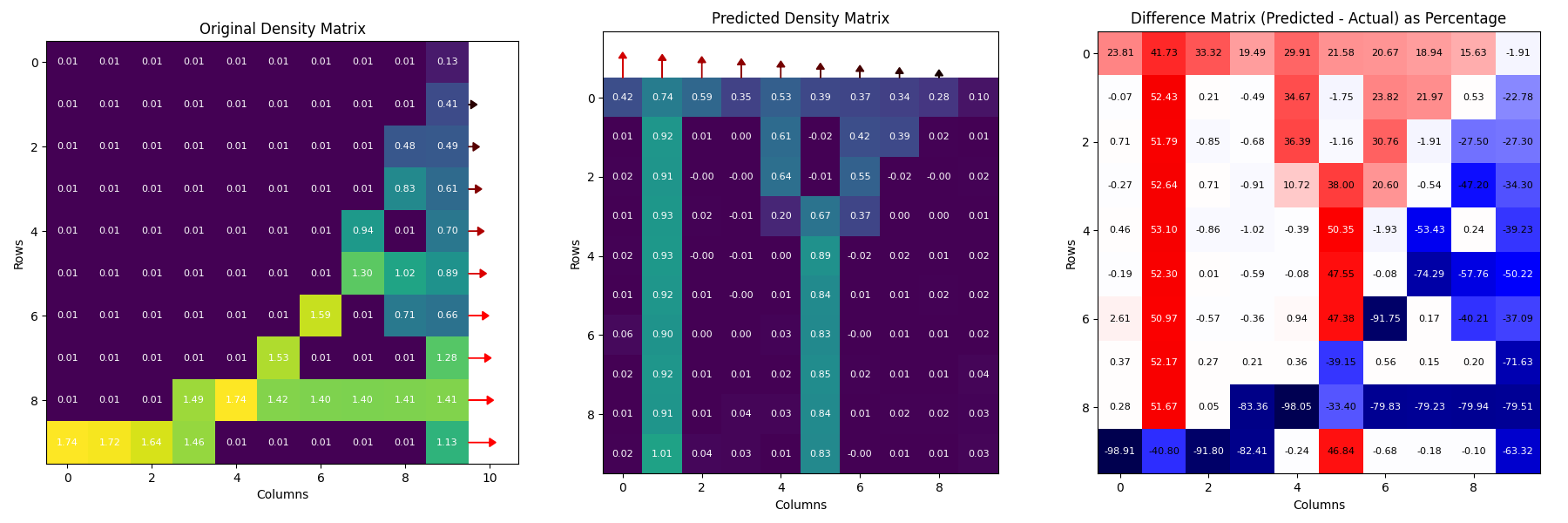
Figure 3: A baseline supervised learning model struggles to solve the inverse bone remodelling model
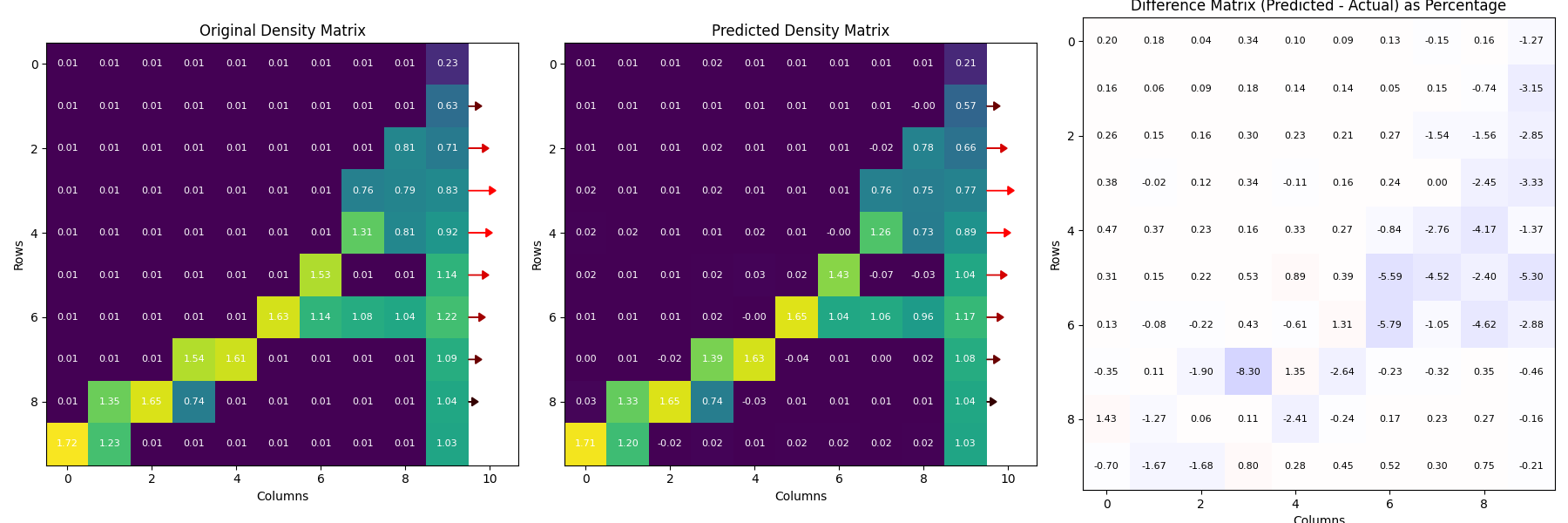
Figure 4: A reinforcement learning framework can estimate the inverse bone remodelling relation
The goal of this thesis is to explore how reinforcement learning can be applied to solve the inverse bone remodelling problem. This repository includes:
- Implementation of reinforcement learning algorithms.
- Simulation environments for testing and evaluation.
- Data preprocessing and analysis scripts.
- Visualisation tools for results and findings.
-
Clone the repository:
git clone https://github.com/gijsschlief/thesis-code.git cd thesis-code -
Install the required dependencies:
pip install poetry
-
Run poetry:
poetry install
(If the poetry install does not work rebuilt the poetry.lock file using: "poetry build" and retry)
-
Install conda (as Fenics can not be installed through Poetry): https://www.anaconda.com/docs/getting-started/miniconda/install#linux-terminal-installer
-
Install Fenics using conda:
conda install -c conda-forge fenics=2019.1.0
- Build:
docker build -t bone-rl-image . - Run:
docker run --rm -v $(pwd)/data:/app/data bone-rl-image
Please be aware that all code has currently been implemented using absolute paths that point to local drives, which need to be changed inside the source code to a new location! The main scripts that contain the important steps taken in the thesis are the files:
- density_animation.py # Run the forward simulation on a sample and plot the animation
- generator.py # Run the forward simulation in parallel on many generated samples
- trainer.py # Train a surrogate model on the created dataset
- evaluate_surrogate.py # Evaluate surrogate performance
- run_rl_environment.py # Train the rl_agent
- evaluate_agent.py # Analyse agent performance on the test set
- inverse_trainer.py # Run the inverse neural network on the same dataset
INVERSE-BONE-REMODELING-RL/
├── delft_blue/ # .sh files for running modules on DelftBlue supercomputer
├── bone_remodelling/ # All source code
│ ├── analysis/ # Analysis tools and plots
│ ├── forward_data/ # Data generation and visualisation
│ ├── forward_model/ # Simulation environment (Forward Model)
│ ├── inverse_model/ # Simulation environment (Inverse Baseline)
│ ├── rl_model/ # RL models and PPO training scripts
│ ├── surrogate_model/# Neural network to estimate the forward model
│ └── main.py # Entry point for running experiments
├── tests/ # Test functions for all modules
└── README.md # Project documentation
Contributions are welcome! Feel free to open issues or submit pull requests to improve the codebase.
This project is licensed under the MIT License. See the LICENSE file for details.
Feel free to reach out if you have any questions or need further assistance!