-
Notifications
You must be signed in to change notification settings - Fork 10
Round 1
As a means to improve upon the initial model created by Murray, OSM sought to utilise the expertise of the scientific community by creating a competition in which anybody could participate in developing a predictive model for the series. The competition was launched in August 2016 and ran until March 2017. During this time, participants were provided with a number of datasets for which to train their models. This included lists of molecular identifiers (e.g. SMILES strings) for active and inactive compounds (OSM, Malaria Box and Maybridge) which had been evaluated in the Kirk ion regulation assay, published data on compounds from the spiroindolone, dihydroisoquinolone and pyrazoleamide structure classes, and the complete list of OSM compounds from Series 1 to 4. In addition to this, the original homology model developed by Jeremy Horst was provided, along with information regarding the relevant mutations known to be associated with resistance to ATP4.
By the end of the competition, six entrants had developed and submitted models. Details of each entrant's model are provided below, along with their submitted entries and ELNs. Test set B was the Frontrunner compounds. Test set C was a list of new Series 4 compounds.
| Entrant | Summary of Model | |
|---|---|---|
| Vito Spadavecchio | Created a library of 'common' transformations as seen in the CHEMBL database of compounds (chembl_22_1), which amounts to ~1.67 million compounds. After all of the SMIRKS/SMARTS patterns were extracted for all of the transformations, I was able to enumerate a new library of compounds based off of (what seems to be) a lead compound for series 4: MMV669844. Next, this library was then applied to my MLP potency prediction algorithm (described ELN); here, I used 'modelB.csv' as my model, with the notable exception that all compounds in test set 'B' were labeled 'A', and thus included in on the training. The new compounds potencies were then predicted; all compounds with sub micromolar potency (e.g. pIC50 < 0) were then selected, and filtered for compounds with certain properties. | Submission ELN |
| Ho-Leung Ng | Tried to predict a binding site and the best fitting ligands: Used Cresset Forge to generate conformers and align with docking model of highly potent compounds, MMV670947, bound to homology model of PfATPase, pose #5, as described in my other notebook. That pose was used as the "Reference" for Forge. 3D alignments are generated with training set composed of Series 4 compounds, excluding those marked as Test Sets B and C, and those with log potency <= 5. The low potency compounds were excluded because assays did not provide precise values, only marking them "< 5", for example. Test Set C was used as the "Prediction Set" by Forge. | Submission ELN |
| Giovanni Cincilla | Developed several PfATP4 Ion Regulation Activity classification models using different strategies for modeling set sampling, different machine learning methods and different descriptors. Submitted the best performing one with which we achieved good general results: balanced accuracy (for actives) = 0.77, sensitivity (for actives) = 0.833, AUC (for actives) = 0.810. | Submission ELN |
| James McCulloch | The final model is a meta classifier which uses the probability maps of upstream classifiers to produce an optimal composite classification. Each predictive model based on fingerprints or another SMILE based description vector such as DRAGON* brings a certain amount of predictive power to the task of assessing likely molecular activity against PfATP4. The meta classifier combines the predictive power of each model in an optimal way to produce a more predictive composite model. It does this by taking as it's input the probability maps (the outputs) of other classifiers. *A Neural Network model that uses the DRAGON molecular descriptor to estimate molecular PfATP4 ion regulation activity directly. This model had modest predictive power of AUC=0.77. **A logistic classifier that uses the Morgan fingerprints (mol radius = 5) to predict the EC50 <= 500 nMol class. This model has a predictive power of AUC=0.93 for the test molecules. | Submission ELN |
| Davy Guan | A semi-supervised machine learning paradigm adapted from the machine learning algorithms implemented in the DeepChem project was used to construct QSAR models from both the labelled and unlabelled datasets. All molecules were featurised by either Graph convolutional techniques or with 1024 Bit ECFP4 descriptors. A 80/10/10 train, test, internal validation was used to split the Training dataset for model construction and internal validation before testing on the external validation dataset. | Submission ELN |
| Johnathan Silva | Gradient boosting model (using xgboost) to predict actives and nonactives for the PfATP4 ion regulation assay. Data sampled to include only those in the vicinity of OSM S4 compounds. | Submission ELN |
The end goal was for the entrants to use their models to rank the compounds from the MMV Pathogen Box from most to least active. The top twenty active compounds from these rankings were evaluated by a judging panel, who compared the predicted compounds against unreleased experimental data obtained by the Kirk lab. In doing so, two models (created by Ho-Leung Ng and James McCulloch) were identified to have correctly predicted two active compounds in their top twenty rankings and one active compound just outside their top twenty (Figure 1). The competition concluded with a tie for first place and both participants were awarded $500.
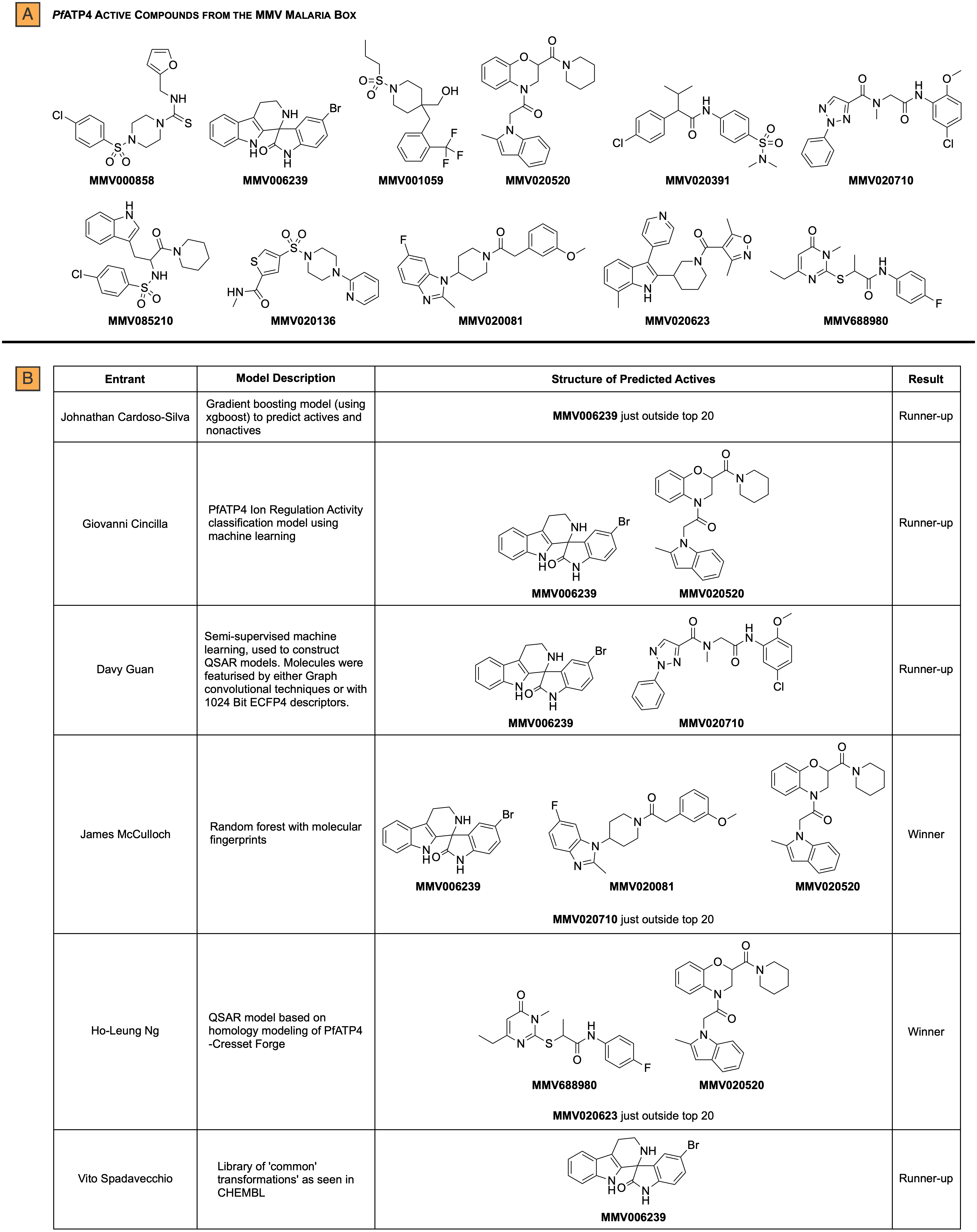
Figure 1: Summary of correct top 20 rankings from each entrant. (A) 11 PfATP4 active compounds from the MMV Malaria Box. (B) Structures of the correctly predicted actives.
In a similar manner to Murray's original model, while the ones created during this competition were able to predict a small number of active compounds correctly, the models were still not highly predictive. Encouragingly, the success of the competition was demonstrated with six new solutions from the community that were based on different approaches, with all entrants working openly and all data being shared.